When you talk about BioClaw, which project are you talking about?
If someone says "have you seen BioClaw?" in a conversation about AI for science, you now need to ask a follow-up question. Because there are two of them.
One was built by a team connected to Princeton University. The other came out of Beijing 1st Biotech Group and the PLA General Hospital. They were developed on opposite sides of the Pacific, apparently without either team knowing the other existed. They do completely different things. And their names are, for all practical purposes, identical.
Welcome to the OpenClaw naming crisis.
The Quick Version
| BioClaw | BloClaw | |
|---|---|---|
| Team | Princeton University (Zaixi Zhang et al.) | Beijing 1st Biotech + PLA General Hospital |
| What it does | Bioinformatics research assistant | Multi-modal scientific workspace |
| Core tools | PubMed search + PyMOL visualization | RDKit + ESMFold + molecular docking + RAG |
| Built on | OpenClaw | Independent |
| Language | TypeScript | Python |
| Paper | bioRxiv (STELLA) | arXiv:2604.00550 |
| GitHub Stars | ~318 | ~1 (just released) |
| Website | bioclaw.tech | — |
Two projects. Two papers. Two completely different approaches to AI-assisted science. One name.
BioClaw: The Focused Bioinformatics Assistant
BioClaw comes from the team behind STELLA (Self-Evolving LLM Agent for Biomedical Research), with Zaixi Zhang at Princeton as the corresponding author. It's built on top of OpenClaw — meaning it inherits the full OpenClaw messaging infrastructure — and focuses on doing two things really well:
- PubMed literature search — query, retrieve, and synthesize biomedical papers directly from within the agent
- PyMOL molecular visualization — generate publication-quality molecular graphics through natural language
It also ships with ~25 built-in skills covering BLAST search, differential expression analysis, single-cell preprocessing, and database queries. There's a web UI with a "Lab Trace" dashboard for tracking what the agent did and why.
The philosophy is clear: don't try to do everything. Do bioinformatics well. If you're a wet-lab biologist who needs to quickly search literature and visualize a protein structure without leaving your chat window, this is your tool.
TypeScript codebase, 318 stars, active development, official project page with demos.
BloClaw: The Omniscient Workspace
BloClaw — and yes, the paper literally uses the word "omniscient" in the title — takes the opposite approach. Where BioClaw is a scalpel, BloClaw wants to be the entire operating room.
Published as "BloClaw: An Omniscient, Multi-Modal Agentic Workspace for Next-Generation Scientific Discovery" on arXiv, this project from Beijing 1st Biotech and the PLA General Hospital covers:
- Cheminformatics via RDKit — molecular property calculation, substructure search, chemical similarity
- Protein structure prediction via ESMFold — de novo 3D folding without MSA
- Molecular docking — predict how small molecules bind to protein targets
- Autonomous RAG — retrieval-augmented generation for scientific literature
The technical architecture is interesting. They built an XML-Regex dual-track routing protocol that eliminates the serialization errors that plague most agent tool-calling systems. There's a runtime state interception sandbox that uses Python monkey-patching to capture data visualizations as they're generated. And the UI is "state-driven dynamic viewport" — meaning it adapts to what the agent is currently doing.
Python codebase, just 1 star (very new), but backed by an arXiv paper with detailed benchmarks.
The Spelling Situation
Here's where it gets genuinely funny.
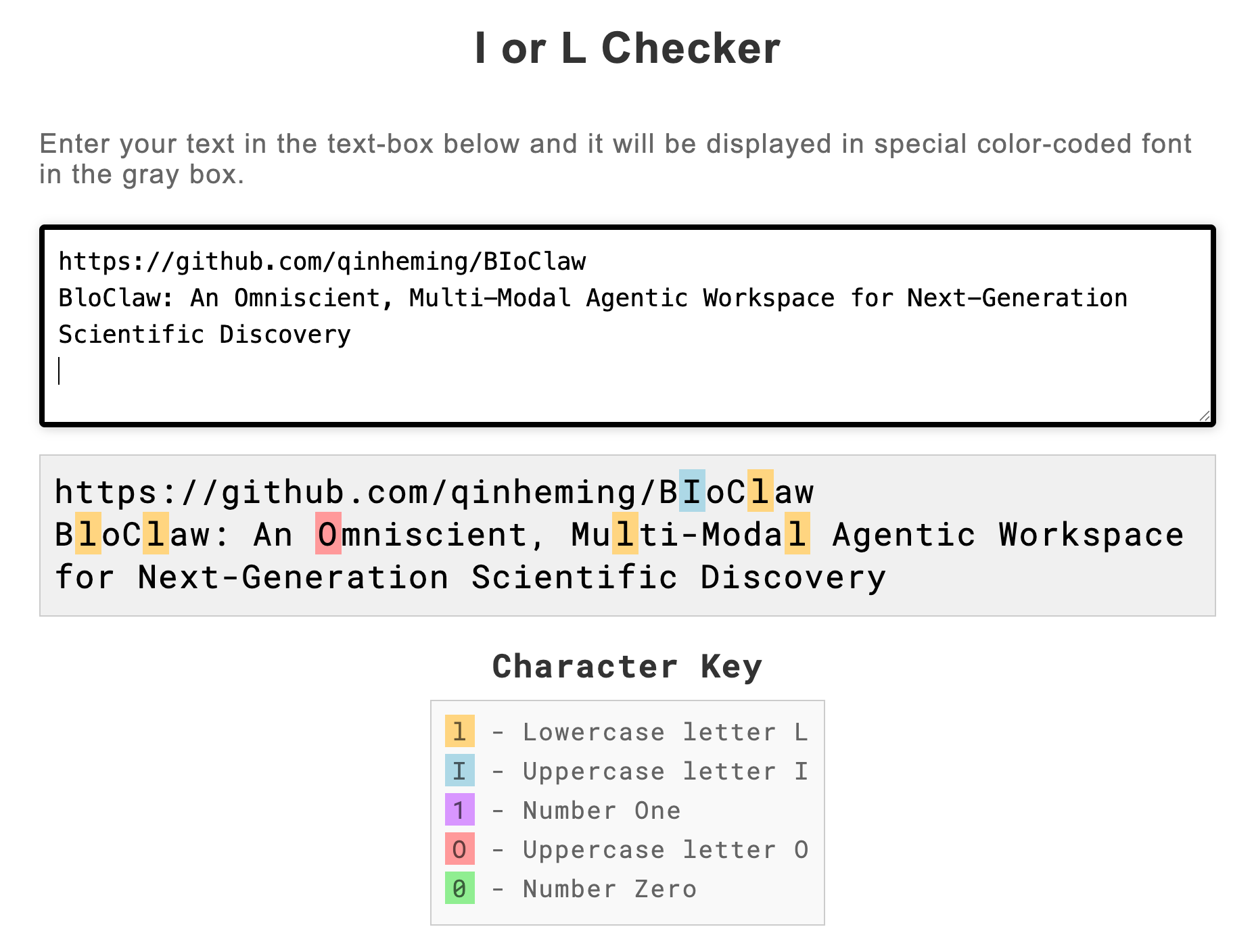
The Beijing team's paper title spells it BloClaw — B, lowercase L, o, C, l, a, w. But their GitHub repository is named BIoClaw — B, uppercase I, o, C, l, a, w.
In most fonts, lowercase L and uppercase I are indistinguishable. The authors accidentally created a project whose name is spelled differently in their own paper versus their own repository, and you literally cannot tell by looking.
We verified this using an I-or-L checker tool that renders each character in a special font. The paper says l. The repo says I. They look identical.
This isn't an isolated incident. The OpenClaw ecosystem now has 9 groups of name collisions we track:
- ScienceClaw (4 projects)
- PaperClaw (6 projects)
- ResearchClaw (4 projects)
- MedClaw (3 projects)
- DrugClaw (2 projects)
- SciClaw (2 projects)
- ClawSafety (2 projects)
- ClawTeam (2 projects, one a fork)
- BioClaw/BloClaw (2 projects)
When an ecosystem grows from zero to 130+ projects in four months with no central naming registry, this is what happens. It's the domain name squatting problem all over again, except nobody's squatting — they just didn't check.
Which One Should You Use?
Choose BioClaw if:
- You're a bioinformatics researcher or wet-lab biologist
- You primarily need literature search and molecular visualization
- You want something that works within the OpenClaw ecosystem
- You value a mature project with demos and documentation
Choose BloClaw if:
- You're in computational chemistry or drug discovery
- You need protein structure prediction and molecular docking
- You want a self-contained workspace (not OpenClaw-dependent)
- You're interested in the latest research (arXiv paper with benchmarks)
Choose neither if:
- You need single-cell RNA-seq analysis → try OmicsClaw
- You need general research automation → try AutoResearchClaw
- You need agent safety evaluation → try ClawSafety or Claw-Eval
The Bigger Picture
Name collisions are annoying, but they're actually a sign of health. Nobody fights over names in a dead ecosystem. The fact that two independent teams — one at an Ivy League university, one at a Chinese biotech company — both decided to build AI science agents using the same naming convention tells you something about where the field is heading.
Every research domain is getting its own agent. Bioinformatics, drug discovery, materials science, clinical research — each one needs specialized tools, specialized knowledge bases, and specialized workflows. The generic "ask ChatGPT to help with your research" era is ending. The specialized agent era is beginning.
And when it's fully here, you'll need a directory to keep track of which agent does what.
That's what we're building.
For a detailed side-by-side comparison, see our BioClaw / BloClaw disambiguation page. Both projects are listed in our project directory — BioClaw under Biomedicine, BloClaw under Drug Discovery.
